Chapter 4: Estimating Copy Number in ddPCR
Digital PCR (ddPCR) partitions a reaction mix into thousands of droplets, each acting as an independent PCR microreaction.
The goal is to estimate the absolute concentration of the target gene in the original sample, based on how many droplets are positive (i.e., contain at least one copy of the target sequence).
Key Parameters
- \(N\): total number of droplets
- \(p\): number of positive droplets (detected signal)
- \(v\): volume of a single droplet (in μL)
- \(d\): dilution factor (from original sample to the PCR well)
Estimating Copy Per Droplet (CPD)
From Poisson theory, the probability that a droplet is negative (i.e., contains no target) is:
\[ P(X = 0) = e^{-\lambda} \]
Rearranging:
\[ \lambda = -\ln\left(\frac{N - p}{N}\right) = -\ln\left(1 - \frac{p}{N}\right) \]
This \(\lambda\) is the mean copy number per droplet.
Estimating Concentration (copies/μL)
We can convert copy per droplet (CPD) into concentration using the droplet volume \(v\) and dilution factor \(d\):
\[ C = \frac{\lambda \cdot d}{v} = -\frac{d}{v} \ln\left(1 - \frac{p}{N}\right) \]
This gives the absolute concentration (copies/μL) in the original sample.
Example 1: Moderate Concentration
- Reaction volume: 20 μL
- Total droplets: 20,000
- Positive droplets: 200
- No dilution (\(d = 1\))
N <- 20000
p <- 200
v <- 20 / N
d <- 1
lambda <- -log(1 - p / N)
C <- lambda * d / v
paste(round(C, 3), "copies/uL")## [1] "10.05 copies/uL"Example 2: Rare Target Detection
Only 1 positive droplet observed among 20,000:
## [1] "0.05 copies/uL"Visualizing Positive Droplets vs. Concentration
library(ggplot2)
result <- data.frame(p.drop = 1:20000)
result$conc <- round(-1 / (20 / 20000) * log(1 - result$p.drop / 20000), 3)
ggplot(result, aes(x = p.drop, y = conc)) +
geom_point(size = 0.1) +
labs(title = "Estimated Concentration vs. Positive Droplets",
x = "Positive Droplets (p)",
y = "Estimated Concentration (copies/uL)")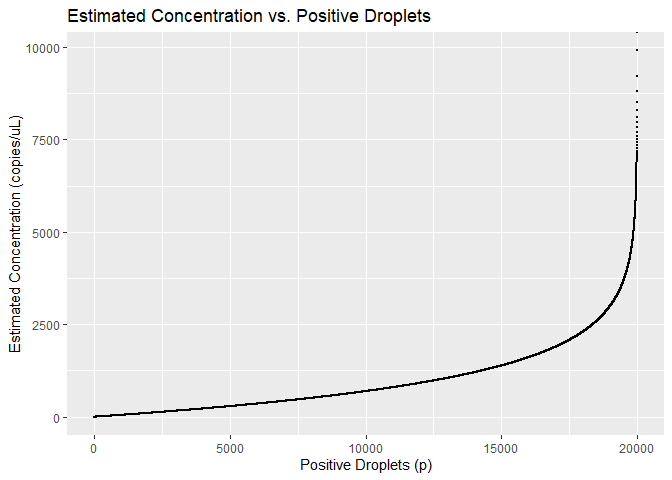
Figure 2: Estimiated concentration vs positive droplet
Summary
- ddPCR uses the fraction of positive droplets to estimate \(\lambda\)
- \(\lambda\) is then converted to copies per μL
- This method requires no standard curve, unlike qPCR
- Particularly useful for low copy number detection and absolute quantification
In the next chapter, we will explore how \(\lambda\) relates to detection probability and how it informs Limit of Detection (LoD) and Confidence Intervals in ddPCR.